The breathtaking diversity of life that surrounds us is the underlying inspiration for our lab’s research and training goals that are powered by collaborations worldwide.
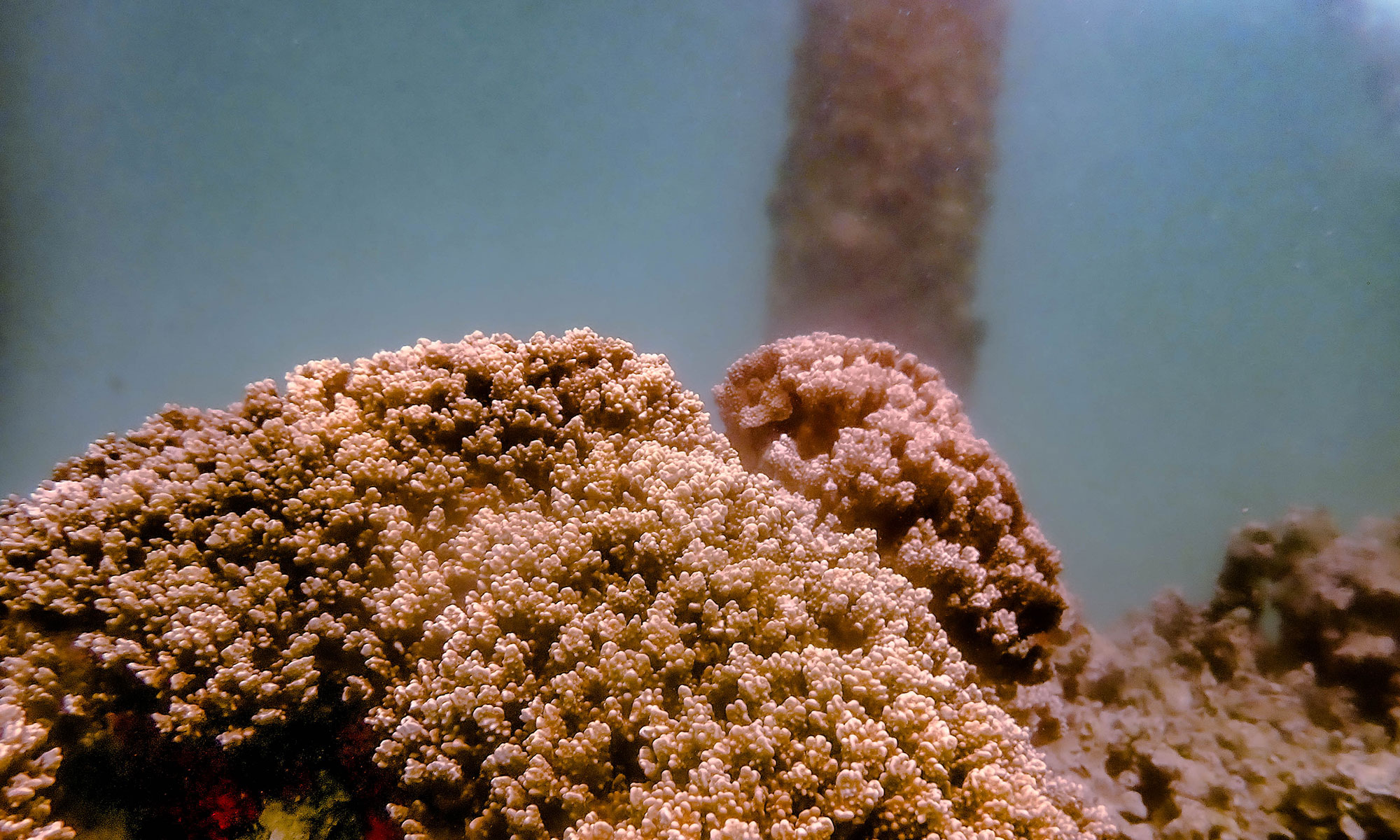
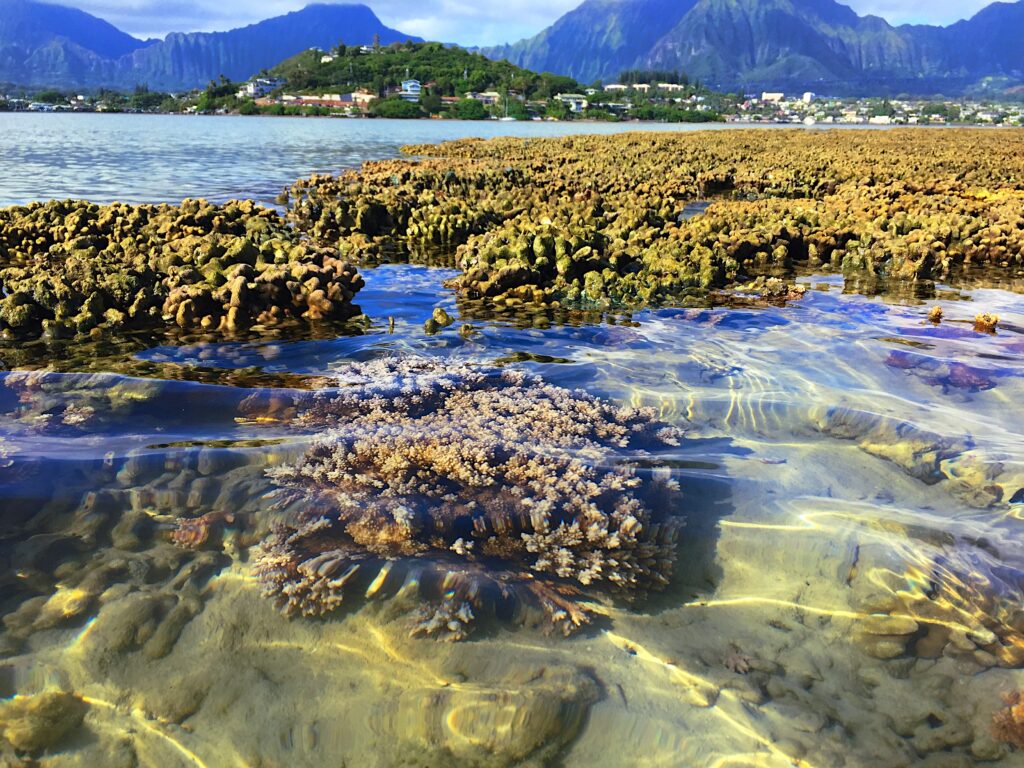
The Bhattacharya Lab (dblab) has long-standing interests in algal/protist and coral evolution and genomics and specific projects include elucidating the endosymbiotic origin of photosynthetic organelles (plastids), advancing coral biology and conservation, erecting and testing the eukaryotic tree of life, understanding the role of horizontal gene transfer in eukaryote evolution, and developing engineering solutions for field monitoring of threatened species.
Latest News
- Work done by Erin Chille and Timothy Stephens in Ulithi Atoll
- Jiayi Pu presents her research poster a the Aresty Undergraduate Research Symposium
- Daisy Cifuentes presents her research poster a the Aresty Undergraduate Research Symposium
- Shriya Singaraju (‘25) was awarded the 2025 D.O.O.R.S. Scholarship (D.O.O.R.S. – Diversification Of Our Research Scientists)
- Pets of Bhattacharya Lab
- Video Release: The Endosymbiotic Ratchet
- Debashish meets with private foundations in Florida and has a close encounter with an alligator.
- Debashish gives invited talk in NYU Abu Dhabi about coral research.
- PHD student Arwa Gabr successfully defends her doctoral thesis.
- Learn about all of the new findings about Paulinella and plastid endosymbiosis
- Bhattacharya lab collaborates with the Kewalo lab in Hawaii to study coral biology.
- Graduate student Shrinivas Nandi joins the Bhattacharya lab.
2024 Papers from the Bhattacharya Lab and Collaborators
Fpa (YlaN) is an iron(II) binding protein that functions to relieve Fur-mediated repression of gene expression in Staphylococcus aureus. 2024. Jeffrey M. Boyd, Kylie Ryan Kaler, Karla Esquilín-Lebrón, Ashley Pall, Courtney J. Campbell, Mary E. Foley, Gustavo Rios-Delgado, Emilee M. Mustor, Timothy G. Stephens, Hannah Bovermann, Todd M. Greco, Ileana M. Cristea, Valerie J. Carabetta, William N. Beavers, Debashish Bhattacharya, Eric P. Skaar, Lindsey N. Shaw, Timothy L. Stemmler. mBio 15:e02310-24.
Biomarkers of Marine Invertebrate Stress
D Bhattacharya, TG Stephens, S Nandi, EE Chille US Patent App. 18/618,557
What Has Paulinella Taught us About Endosymbiont Metabolic Integration? 2024. V Calatrava, TG Stephens, AR Grossman, D Bhattacharya. In: Schwartzbach, S.D., Kroth, P.G., Oborník, M. (eds) Endosymbiotic Organelle Acquisition. Springer, Cham.
The Evolutionary Origin of Primary Plastids. 2024. D Lhee, D Bhattacharya, HS Yoon. In: Schwartzbach, S.D., Kroth, P.G., Oborník, M. (eds) Endosymbiotic Organelle Acquisition. Springer, Cham.
Gene expression response under thermal stress in two Hawaiian corals is dominated by ploidy and genotype. 2024. Erin E. Chille, Timothy G. Stephens, Deeksha Misri, Emma L. Strand, Hollie M. Putnam, Debashish Bhattacharya. Ecology and Evolution, 14, e70037.
Evidence of a putative CO2 delivery system to the chromatophore in the photosynthetic amoeba Paulinella. 2024. Arwa Gabr, Timothy G. Stephens, John R. Reinfelder, Pinky Liau, Victoria Calatrava, Arthur R. Grossman, Debashish Bhattacharya. Environmental Microbiology Reports, 16(3), e13304.
Whole-genome duplication in an algal symbiont bolsters coral heat tolerance. 2024. Katherine E. Dougan, Anthony J. Bellantuono, Tim Kahlke, Raffaela M. Abbriano, Yibi Chen, Sarah Shah, Camila Granados-Cifuentes, Madeleine J. H. Van Oppen, Debashish Bhattacharya, David J. Suggett, Mauricio Rodriguez-Lanetty , and Cheong Xin Chan. Science Advances.
From dusk till dawn: cell cycle progression in the red seaweed Gracilariopsis chorda (Rhodophyta). 2024. JM Lee, S Miyagishima, D Bhattacharya, HS Yoon. iScience.
Nuclear genomes of dinoflagellates reveal evolutionarily conserved pattern of RNA editing relative to stress response. 2024. Y Chen, KE Dougan, D Bhattacharya, CX Chan. Frontiers in Protistology 2, 1320917.
Diverse fates of ancient horizontal gene transfers in extremophilic red algae. 2024. J Van Etten, TG Stephens, E Chille, A Lipzen, D Peterson, K Barry, …Environmental microbiology 26 (5), e16629.
Hot springs viruses at Yellowstone National Park have ancient origins and are adapted to thermophilic hosts. 2024. L Felipe Benites, TG Stephens, J Van Etten, T James, WC Christian, …Communications Biology 7 (1), 312.
Genome-wide transcriptome analysis reveals the diversity and function of long non-coding RNAs in dinoflagellates. 2024. Y Chen, KE Dougan, Q Nguyen, D Bhattacharya, CX Chan. NAR Genomics and Bioinformatics 6 (1), lqae016
Exceptional Quantum Efficiency Powers Biomass Production in Halotolerant Algae Picochlorum sp.^. 2024. C Gates, G Ananyev, F Foflonker, D Bhattacharya, GC Dismukes. Photosynthesis Research, 1-19.
Diurnal Rhythms in the Red Seaweed Gracilariopsis chorda are Characterized by Unique Regulatory Networks of Carbon Metabolism. 2024. JM Lee, JH Yang, APM Weber, D Bhattacharya, WY Kim, HS Yoon. Molecular Biology and Evolution 41 (2), msae012.
Massive genome reduction predates the divergence of Symbiodiniaceae dinoflagellates. 2024. S Shah, KE Dougan, Y Chen, R Lo, G Laird, MDA Fortuin, SK Rai, … The ISME Journal 18 (1), wrae059.